with New England BioLabs - NEBNext Ultra II RNA kits
 |
||||||||||
 |
NEBNext – JUNE 2017 |
|||||||||
 |
||||||||||
| INTRODUCING NEBNEXT ULTRA II FOR RNA | ||||||||||
| Do you need increased sensitivity and specificity from your RNA-seq experiments? Do you have ever-decreasing amounts of input RNA? Our next generation of RNA library prep kits have been reformulated at each step, resulting in several fold higher yields of high quality libraries, enabling use of lower input amounts and fewer PCR cycles. Our new NEBNext® Ultra™ II RNA kits have streamlined, automatable workflows and are available for directional (strand-specific, using the "dUTP method") and non-directional library prep and are compatible with poly(A) mRNA enrichment or rRNA depletion. The kits are available with the option of SPRISelect® beads for size-selection and clean-up steps.
|
||||||||||
|
||||||||||
 |
||||||||||
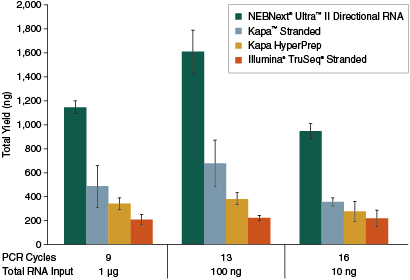
| Poly(A)-containing mRNA was isolated from 10 ng, 100 ng and 1 µg of Universal Human Reference RNA (Agilent® #740000) and libraries were made using the NEBNext Ultra II Directional RNA Kit (plus the NEBNext poly(A) mRNA Magnetic Isolation Kit), Kapa Stranded mRNA-Seq Kit, Kapa mRNA HyperPrep Kit and Illumina TruSeq Stranded mRNA Kit. The input RNA amount and number of PCR cycles are indicated. Library yields from an average of three replicates are shown. | ||||||||||
|
||||||||||
 |
||||||||||
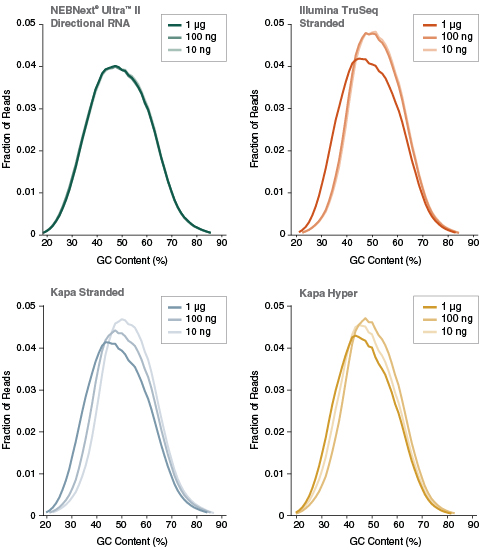
| Poly(A)-containing mRNA was isolated from Universal Human Reference RNA (Agilent #740000), and libraries were made using the NEBNext Ultra II Directional RNA Kit (plus the NEBNext Poly(A) mRNA Magnetic Isolation Module), Illumina TruSeq Stranded mRNA Kit, Kapa Stranded mRNA-Seq Kit and Kapa mRNA HyperPrep Kit. Libraries were sequenced on an Illumina NextSeq® 500 using paired-end mode (2x76 bp). Reads were mapped to the hg19 reference genome. GC content distribution for each library was calculated using mapped reads. Ultra II Directional RNA libraries had uniform GC content distribution across a range of input amounts, whereas for other kits the GC content distribution changed with different input amounts, indicating the introduction of input-dependent sequence bias. | ||||||||||
  |
||||||||||
|
|
||||||||||
 |
||||||||||
|
||||||||||
|
||||||||||












